Returns forecasts and other information for univariate neural network models.
Arguments
- object
An object of class
nnetarresulting from a call tonnetar().- h
Number of periods for forecasting. If
xregis used,his ignored and the number of forecast periods is set to the number of rows ofxreg.- PI
If
TRUE, prediction intervals are produced, otherwise only point forecasts are calculated. IfPIisFALSE, thenlevel,fan,bootstrapandnpathsare all ignored.- level
Confidence levels for prediction intervals.
- fan
If
TRUE,levelis set toseq(51, 99, by = 3). This is suitable for fan plots.- xreg
Future values of any regression variables. A numerical vector or matrix of external regressors; it should not be a data frame.
- lambda
Box-Cox transformation parameter. If
lambda = "auto", then a transformation is automatically selected usingBoxCox.lambda. The transformation is ignored if NULL. Otherwise, data transformed before model is estimated.- bootstrap
If
TRUE, then prediction intervals are produced by simulation using resampled errors (rather than normally distributed errors). Ignored ifinnovis notNULL.- npaths
Number of sample paths used in computing simulated prediction intervals.
- innov
Values to use as innovations for prediction intervals. Must be a matrix with
hrows andnpathscolumns (vectors are coerced into a matrix). If present,bootstrapis ignored.- ...
Additional arguments passed to
simulate.nnetar().
Details
Prediction intervals are calculated through simulations and can be slow. Note that if the network is too complex and overfits the data, the residuals can be arbitrarily small; if used for prediction interval calculations, they could lead to misleadingly small values. It is possible to use out-of-sample residuals to ameliorate this, see examples.
forecast class
An object of class forecast is a list usually containing at least
the following elements:
- model
A list containing information about the fitted model
- method
The name of the forecasting method as a character string
- mean
Point forecasts as a time series
- lower
Lower limits for prediction intervals
- upper
Upper limits for prediction intervals
- level
The confidence values associated with the prediction intervals
- x
The original time series.
- residuals
Residuals from the fitted model. For models with additive errors, the residuals will be x minus the fitted values.
- fitted
Fitted values (one-step forecasts)
The function summary can be used to obtain and print a summary of the
results, while the functions plot and autoplot produce plots of the forecasts and
prediction intervals. The generic accessor functions fitted.values and residuals
extract various useful features from the underlying model.
Examples
## Fit & forecast model
fit <- nnetar(USAccDeaths, size = 2)
fcast <- forecast(fit, h = 20)
plot(fcast)
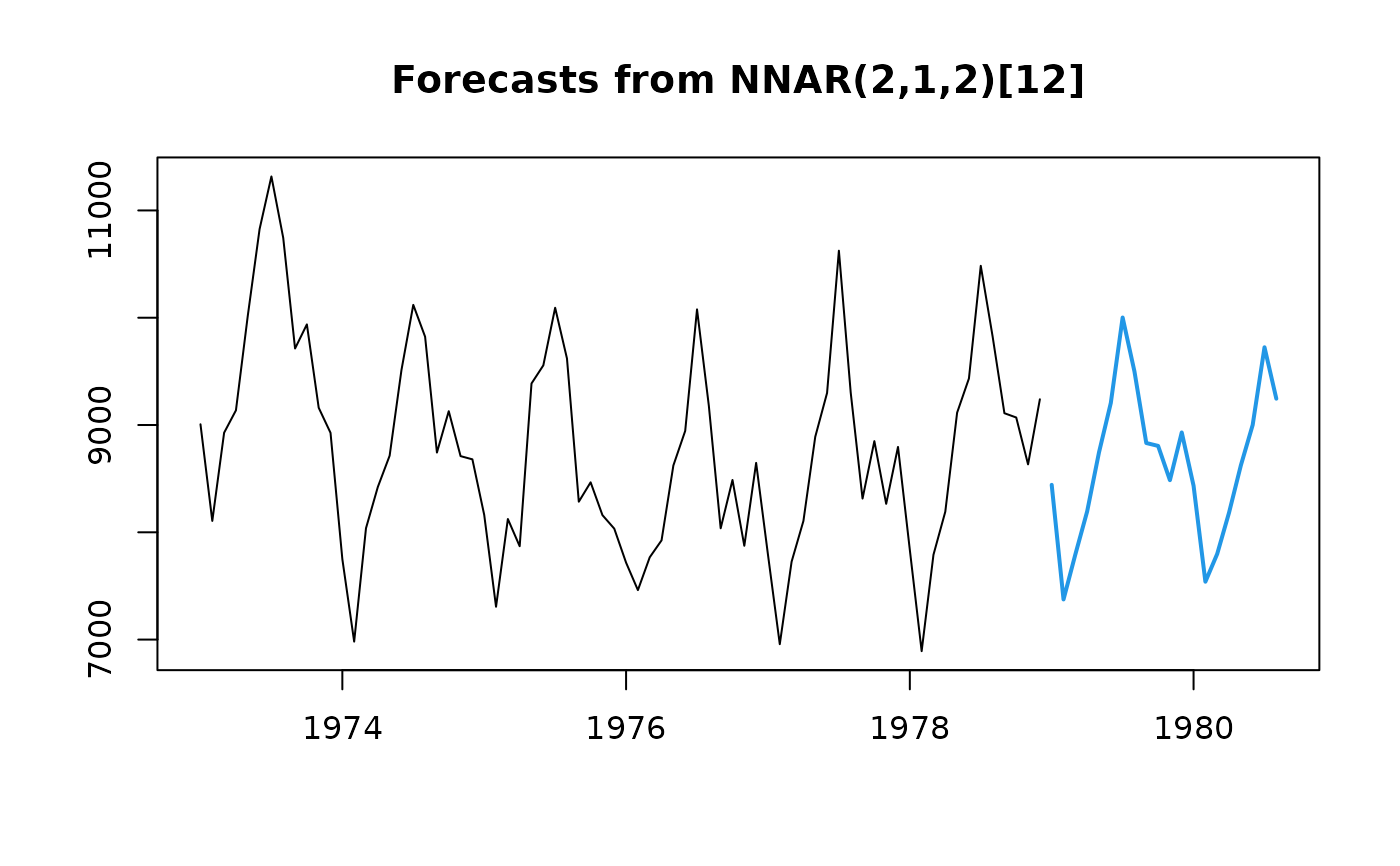 if (FALSE) { # \dontrun{
## Include prediction intervals in forecast
fcast2 <- forecast(fit, h = 20, PI = TRUE, npaths = 100)
plot(fcast2)
## Set up out-of-sample innovations using cross-validation
fit_cv <- CVar(USAccDeaths, size = 2)
res_sd <- sd(fit_cv$residuals, na.rm = TRUE)
myinnovs <- rnorm(20 * 100, mean = 0, sd = res_sd)
## Forecast using new innovations
fcast3 <- forecast(fit, h = 20, PI = TRUE, npaths = 100, innov = myinnovs)
plot(fcast3)
} # }
if (FALSE) { # \dontrun{
## Include prediction intervals in forecast
fcast2 <- forecast(fit, h = 20, PI = TRUE, npaths = 100)
plot(fcast2)
## Set up out-of-sample innovations using cross-validation
fit_cv <- CVar(USAccDeaths, size = 2)
res_sd <- sd(fit_cv$residuals, na.rm = TRUE)
myinnovs <- rnorm(20 * 100, mean = 0, sd = res_sd)
## Forecast using new innovations
fcast3 <- forecast(fit, h = 20, PI = TRUE, npaths = 100, innov = myinnovs)
plot(fcast3)
} # }
