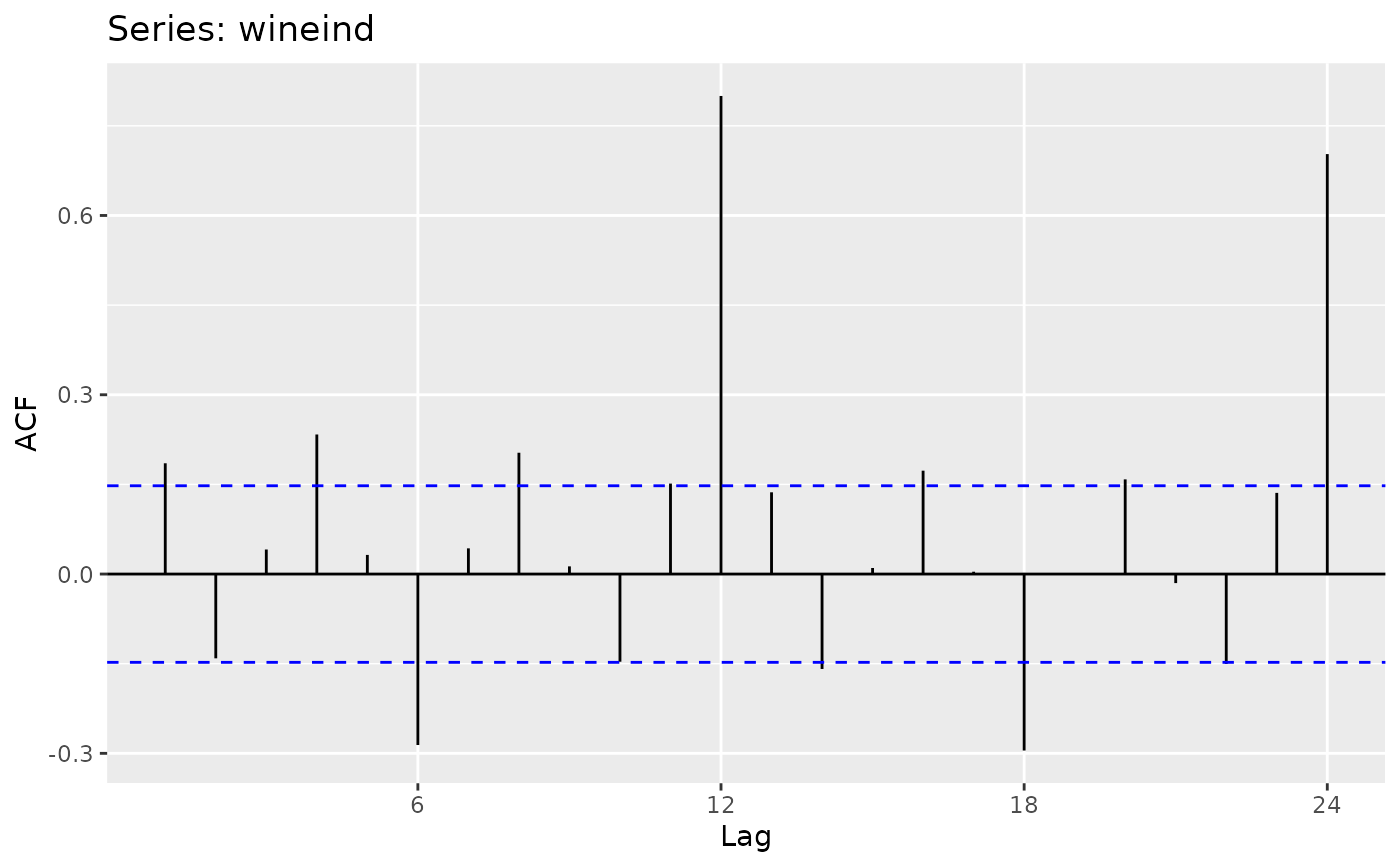
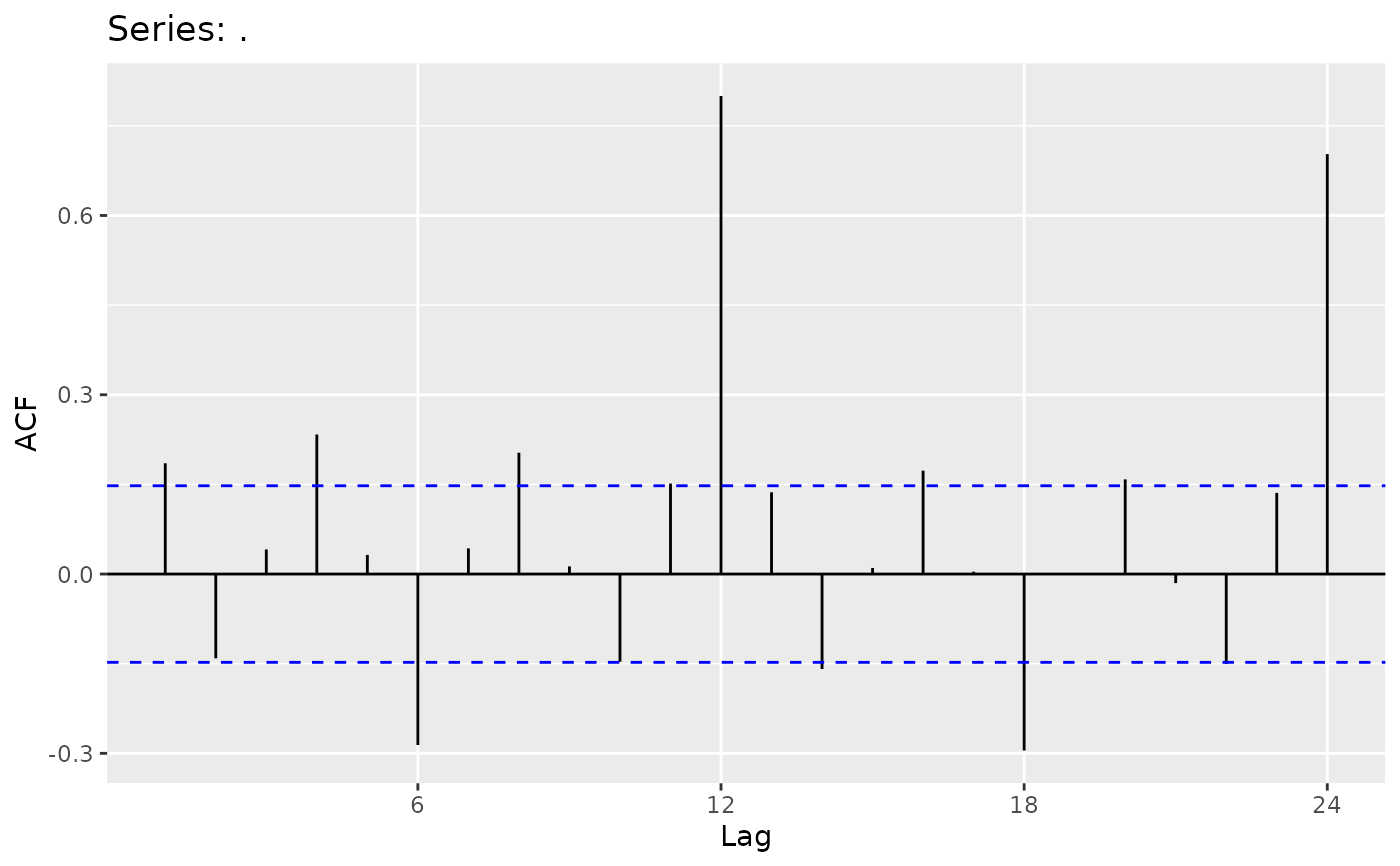
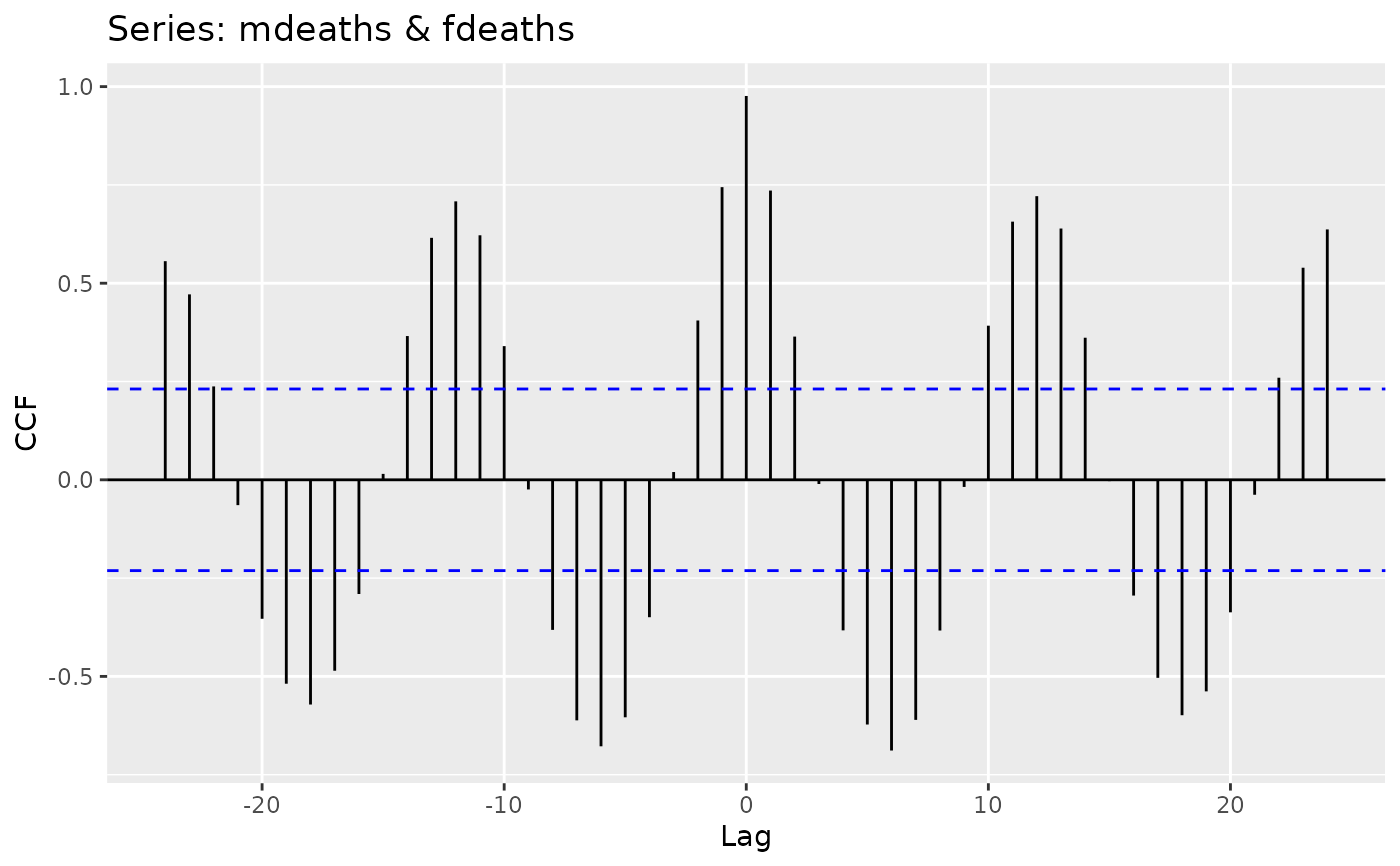
ggplot (Partial) Autocorrelation and Cross-Correlation Function Estimation and Plotting
Source:R/ggplot.R
autoplot.acf.RdProduces a ggplot object of their equivalent Acf, Pacf, Ccf, taperedacf and taperedpacf functions.
Usage
# S3 method for acf
autoplot(object, ci = 0.95, ...)
ggAcf(
x,
lag.max = NULL,
type = c("correlation", "covariance", "partial"),
plot = TRUE,
na.action = na.contiguous,
demean = TRUE,
...
)
ggPacf(
x,
lag.max = NULL,
plot = TRUE,
na.action = na.contiguous,
demean = TRUE,
...
)
ggCcf(
x,
y,
lag.max = NULL,
type = c("correlation", "covariance"),
plot = TRUE,
na.action = na.contiguous,
...
)
# S3 method for mpacf
autoplot(object, ...)
ggtaperedacf(
x,
lag.max = NULL,
type = c("correlation", "partial"),
plot = TRUE,
calc.ci = TRUE,
level = 95,
nsim = 100,
...
)
ggtaperedpacf(x, ...)Arguments
- object
Object of class “
acf”.- ci
coverage probability for confidence interval. Plotting of the confidence interval is suppressed if ci is zero or negative.
- ...
Other plotting parameters to affect the plot.
- x
a univariate or multivariate (not Ccf) numeric time series object or a numeric vector or matrix.
- lag.max
maximum lag at which to calculate the acf.
- type
character string giving the type of acf to be computed. Allowed values are "
correlation" (the default), “covariance” or “partial”.- plot
logical. If
TRUE(the default) the resulting ACF, PACF or CCF is plotted.- na.action
function to handle missing values. Default is
na.contiguous. Useful alternatives arena.passandna.interp.- demean
Should covariances be about the sample means?
- y
a univariate numeric time series object or a numeric vector.
- calc.ci
If
TRUE, confidence intervals for the ACF/PACF estimates are calculated.- level
Percentage level used for the confidence intervals.
- nsim
The number of bootstrap samples used in estimating the confidence intervals.
Details
If autoplot is given an acf or mpacf object, then an
appropriate ggplot object will be created.
ggtaperedpacf
See also
plot.acf, Acf,
acf, taperedacf